Biochemistry and Molecular Biology Committee Faculty
BMB committee members are actively engaged in research projects that represent a diverse range of perspectives. Active student participation is important to their work, and past research students have been co-authors of numerous scholarly publications and presentations at national and international scientific conferences. Students can also work with a variety of other faculty members from allied departments. Current research topics include: the roles of membrane proteins in the regulation of microbial cell growth; design of computer algorithms for analysis of gene and protein structure; mechanisms of gene regulation in animal cells and microorganisms; mechanisms of pathogenesis in herpes viruses; mechanisms of ion channel localization in synaptic neurons; cell cycle regulation in yeast and cancer cells; mechanisms of root gravitropism; identification of bio-active compounds in nature; and computational design of protein-binding drugs.
 Jonathan Fitz Gerald
Jonathan Fitz Gerald
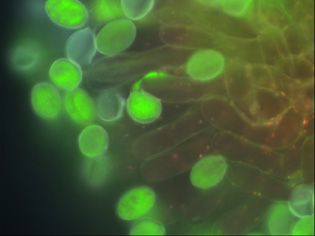
One of the agriculturally significant aspects of plant growth is seed size. This is largely determined by the development of the seed endosperm. Dr. Fitz Gerald′s work focuses on the Arabidopsis gene AtFH5, a formin involved in the development of the posterior endosperm. Using a combination of molecular biology, genetics and microscopy, his aim is to understand both the role of AtFH5 in endosperm development and the pathways that regulate AtFH5 expression. Interestingly, after fertilization AtFH5 is expressed only from the maternal genome. Paternal silencing is regulated by a homologue of the animal Polycomb group complex. In animals, Polycomb complexes maintain cell identity during development. In Arabidopsis, is Polycomb maintaining male and female identity of the parental genomes? Ongoing projects include the examination of mutant plants where AtFH5 expression is altered and molecular screening for AtFH5 interacting proteins. More about Dr. Fitz Gerald′s research
Elaine Frawley
To survive within host organisms and a wide variety of external environments, pathogenic bacteria have evolved flexible metabolic pathways and stress response systems. The Frawley lab studies the physiological, transcriptional and biochemical responses of the model bacterium Salmonella Typhimurium to reactive oxygen and nitrogen species such as hydrogen peroxide and nitric oxide. These host-generated chemical defenses inhibit the function of many key enzymes and so to survive, bacteria must adapt to use alternative enzymes and pathways as well as repair damage that occurs. The lab is currently focused on understanding how nitric oxide stress affects metal homeostasis and the function of metal-containing enzymes by studying the expression and function of bacterial manganese and copper transport proteins.
 Terry Hill
Terry Hill
Dr. Hill’s research deals with the genetic determinants of growth and division in fungal cells. Particular areas of interest are the identification of proteins participating in construction and constriction of actomyosin rings at division sites and the molecular basis for targeting of proteins to their sites of action in the cell. In collaboration with Dr. Loretta Jackson-Hayes (Department of Chemistry), this laboratory is generating and characterizing mutant strains of the filamentous fungus Aspergillus nidulans that have defects in cell growth or division, which is providing information on the mechanisms underlying these processes. Among genes currently under study, are actin, type II myosin, protein kinase C, and a fungal IQGAP protein. Other recently published work has dealt with proteins in the fungal Golgi apparatus, which are important to protein glycosylation and to cell morphogenesis.
Loretta Jackson-Hayes
Research in the Jackson-Hayes lab focuses on investigating regulatory mechanisms of eukaryotic gene expression, focusing on genes that are involved in fungal cell wall metabolism. The fungal cell wall, which is composed of polysaccharides and glycoproteins, is essential for growth and metabolism of the fungus and is an excellent target for antifungal drugs. We have identified several genes that play specific roles in cell wall metabolism in the filamentous fungus Aspergillus nidulansincluding genes that have homologues that have been found to be involved in establishing and maintaining cell wall integrity in yeasts and several other previously uncharacterized genes. Experiments are being conducted to investigate the regulation of message production of these genes, to observe the cellular localization of the proteins during different stages of fungal development, and to learn more about the specific role of each of them in cell wall metabolism.
 Mary Miller
Mary Miller
The growth and division of eukaryotic cells is a highly regulated process. A variety of events important for successful division must be carried out in the proper order, at the proper time, and in the proper location. This coordinated series of events is described as the “cell division cycle” or “cell cycle”. Successful regulation of the cell cycle is paramount to the survival of single and multi-celled organisms ranging from budding yeast to man (Movie of dividing yeast courtesy of M. Tyers). Errors in this process usually result in cell death, and at times trigger the accumulation of oncogenic properties, leading eventually to cancer. In her lab, Dr. Miller studies regulatory proteins called cyclins that trigger coordinated cell division. Dr. Miller′s lab uses the model system Saccharomyces cerevisiae to carry out genetic, genomic, and biochemical assays on cyclin function.
 Larryn Peterson
Larryn Peterson
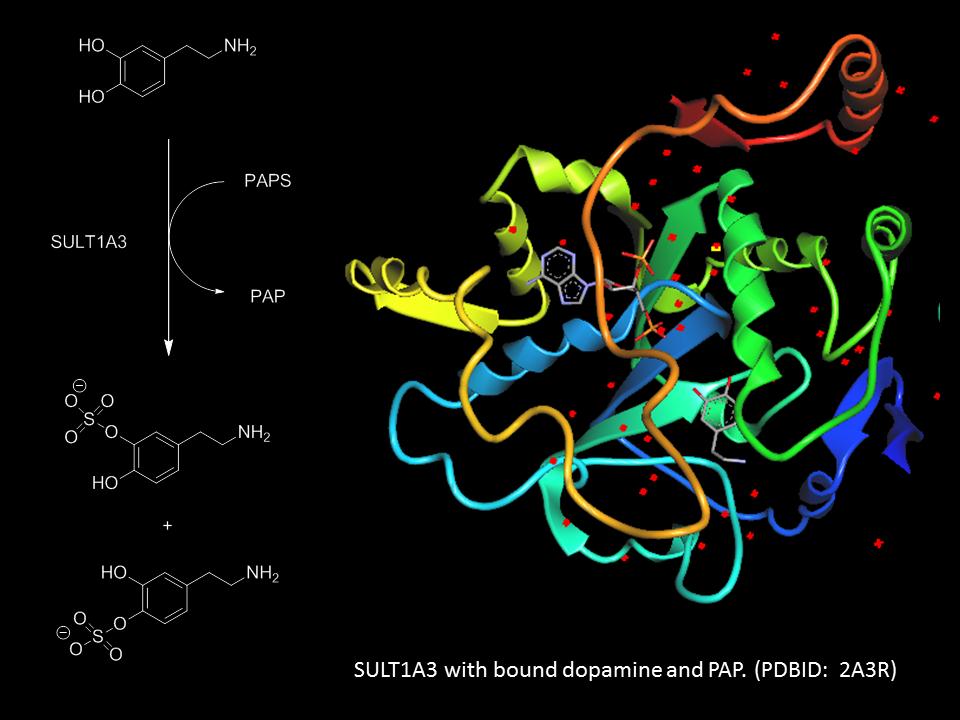
Research in the Peterson lab uses an interdisciplinary approach to understand important biological processes by utilizing tools from synthetic organic chemistry, chemical biology, biochemistry and molecular biology. We currently have two projects ongoing in our lab: 1) investigating the substrate of selectivity of sulfotransferases, a class of enzymes that play a critical role in the regulation and activities of many endogenous molecules including neurotransmitters and hormones, as well as the xenobiotics, and 2) probing the active site requirements of LpxC, an enzyme involved in the biosynthesis of Lipid A, an integral component of Gram-negative bacteria cell walls.
 Dr. Shana Stoddard
Dr. Shana Stoddard
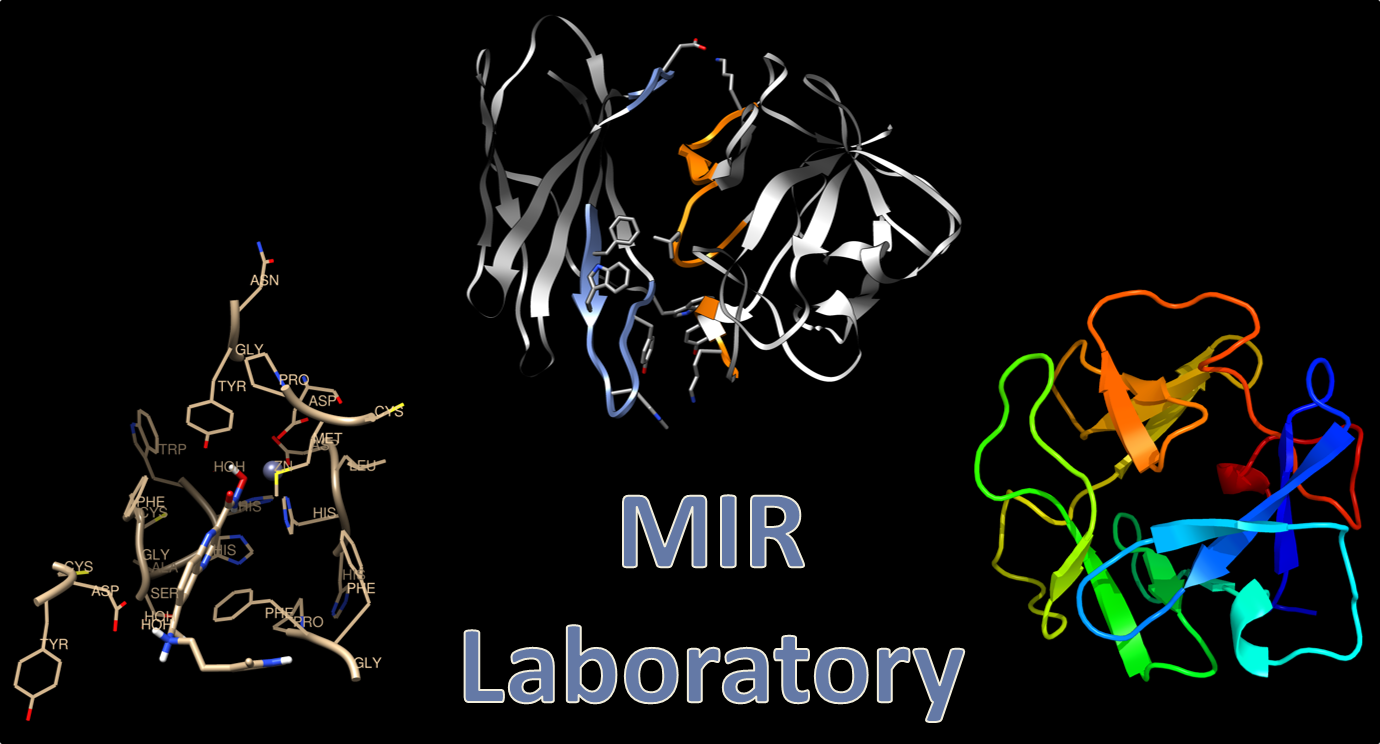
Dr. Stoddard’s research focuses on three aims: (1.) Improving patient outcomes with autoimmune disorders, in particular idiopathic membranous nephritis (IMN), through development of auto-antibody-specific inhibitors, antigen-specific therapies, as well as novel methods for development of antigen specific therapies, (2.) design of novel cancer therapeutics targeting histone deacetylase isozymes, and (3.) understanding the structure and function of antigens involved in autoimmune disorders. In the Molecular Immunotherapeutics Research (MIR) lab we accomplish these goals using a wide range of both computational and experimental research techniques utilized in biochemistry, drug design, and immunochemistry.
Bayly Wheeler
Dr. Wheeler's research is driven by the question of how do cells with identical genomes maintain such diverse patterns of gene expression. In multicellular organisms, cell types with different functions have distinct patterns of gene expression. Remarkably, cells maintain these patterns even after cell division, suggesting that there is a heritable signal that transmits information about gene expression to future generations of cells. One candidate signal is chromatin. The type of chromatin into which DNA is packaged can control gene expression and, if inherited, could allow cells to maintain appropriate patterns of gene expression. Previously, Dr. Wheeler has shown that heterochromatin, a type of chromatin that represses gene expression, can indeed be inherited during cell division in the fission yeast Schizosaccharomyces pombe. This work suggests that heterochromatin may have an important role in maintaining different gene expression patterns among genetically identical cells. Future work in the Wheeler lab will be focused on elucidating the mechanisms that transmit heterochromatin through cell division. Dr. Wheeler is looking forward working with Rhodes students interested in genetics, molecular biology, genomics, epigenetics, and bioinformatics.
